|
Also, indentation in the YAML header has a meaning, so be careful when aligning text. If you need multiple output formats for your R markdown document check whether your YAML options are compatible between these formats. Just a note of caution, many of the options you can specify in the YAML header will work with both HTML and pdf formatted documents, but not all. If you want to explore the plethora of other options see here. You can also change the default font and font size for the whole document and even include fancy options such as a table of contents and inline references and a bibliography. If you would like to change the output to pdf format then you can change it from output: html_document to output: pdf_document (you can also set more than one output format if you like). In the YAML header above the output format is set to HTML. title : My first R markdown document author : Jane Doe date : Maoutput : html_document. 1.4.2 Integrated developement environements.You can toggle between scientific notation preferences in chunks of code by turning it off as described above with options(scipen=999) and reactivating it with options(scipen=0). We can see this scaling work with our big_number variable of 28 billion, reducing it to the expected value of 28.  
Label_number(scale = 1e-9, prefix = "$", suffix = "b", accuracy = 1)) Ggplot(ame(market_cap = caps), aes(x = market_cap)) + 
This tells the computer to scale back the data by 10^-9. Here we call scale_x_continuous and pass the label_number function along with scale = 1e-9 in the labels argument. Thankfully we can use library(scales) to convert the axis labels to something more visually appealing. Is this easier to read? Probably not, even if commas were added to breakup all those zeros. Labs(title = "Market cap for 500 global companies", Geom_histogram(color = "black", fill = "#1FA187") + ggplot(ame(market_cap = caps), aes(x = market_cap)) + options(scipen=999)įormat(summary(caps), big.mark = ",") # using big.mark to add commas # Min.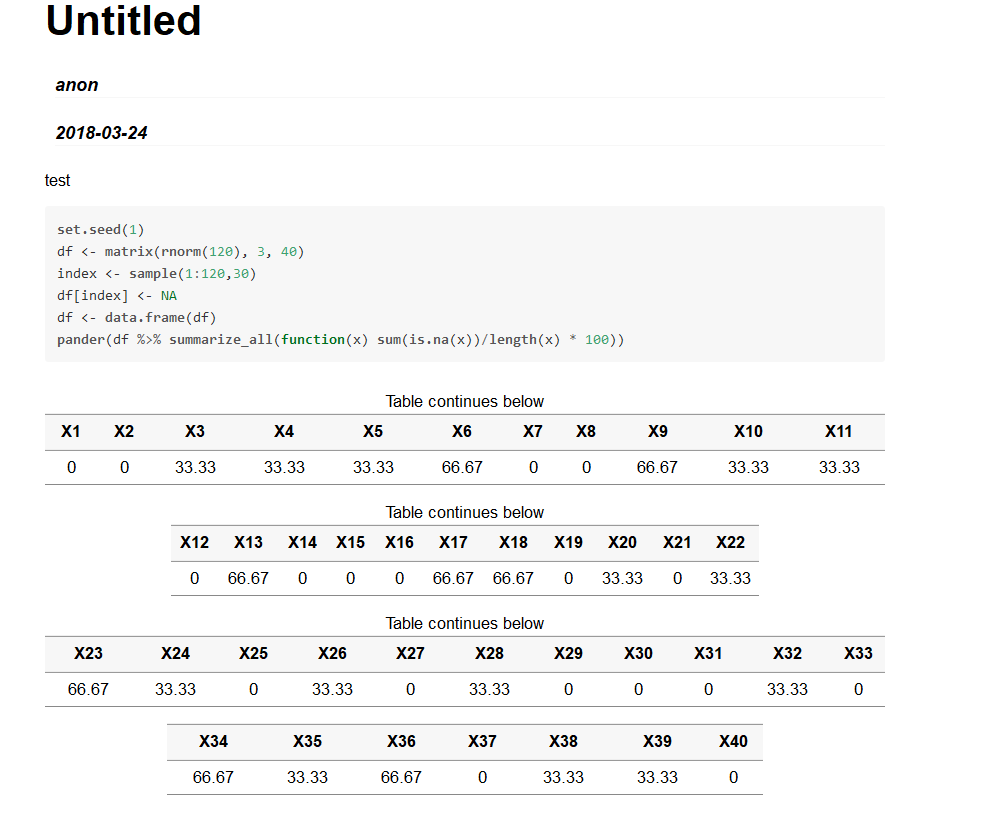 
I generally apply the more aggressive solution by telling R to avoid all scientific notation by setting options(scipen=999) at the top of the script. You can address the newly surfaced visual problem of too many zeros by adding comma separators with big.mark = ',' format(big_number, scientific = FALSE, big.mark = ',') # "28,000,000,000" Universal If you want to avoid scientific notation for a given number or a series of numbers, you can use the format() function by passing scientific = FALSE as an argument. Side-stepping the interpretation problem Targeted Kable(sci, col.names = c("Scientific Notation", "Full digit equivalent")) Our summary statistics in R would look like this: set.seed(123)Ĭaps % mutate(translation = format(sci_note, scientific = FALSE, big.mark = ",")) Imagine we’re dealing with 500 global companies that have an average market cap of 28 billion dollars with a standard deviation of 8 billion. The number of blog posts debating Python vs. R. Sometimes you work with numbers that are pretty big.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
